The Department of Medicine at Cambridge is one of the most prestigious and top ranked in the UK and throughout the world.
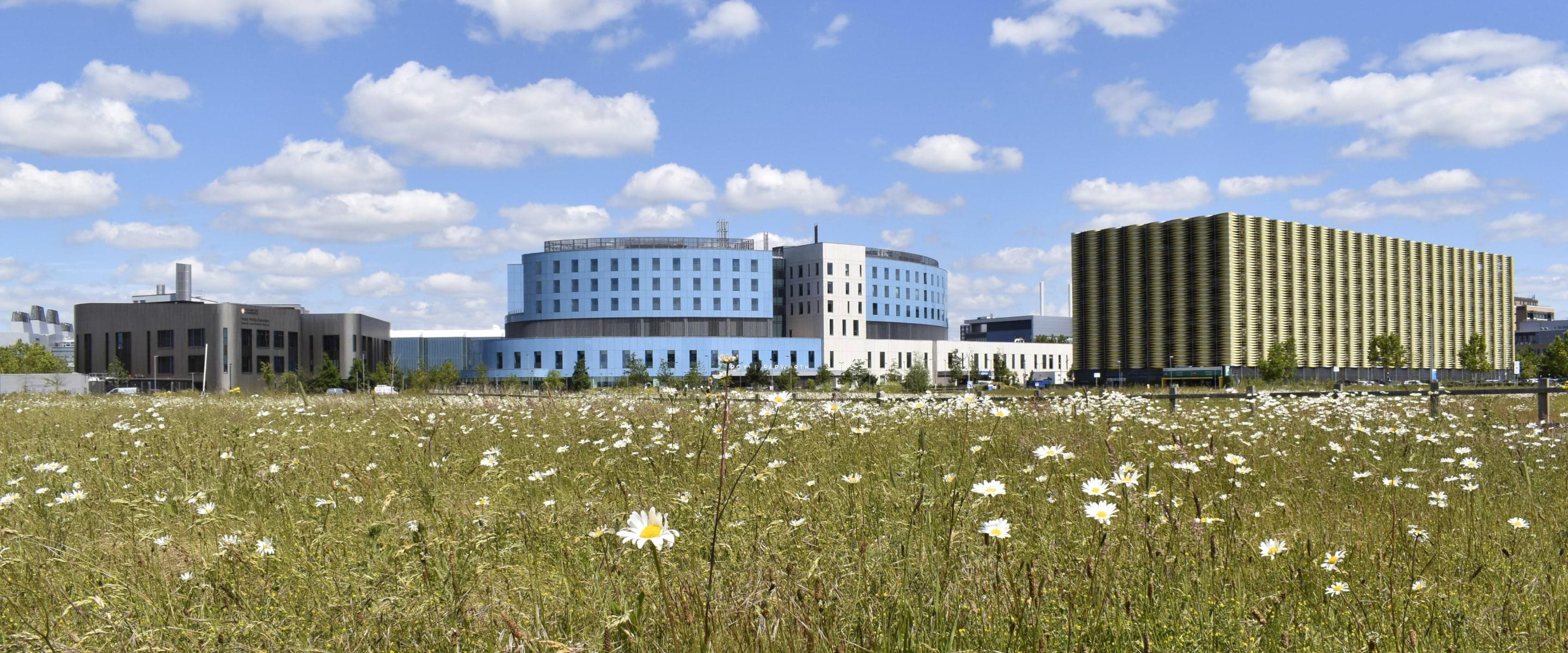
We are the largest department in the School of Clinical Medicine at the University of Cambridge. Our work extends from biomedical research through to the clinic, and is linked to all research throughout the Cambridge Biomedical Campus, the UK, and the wider international community.
Interaction with industry is a major focus of our work, building on the longstanding presence of the GSK Cambridge Clinical Unit, the recent arrival of the Global Research and Corporate Headquarters of AstraZeneca and the growing presence of BioNTech.
750staff, students, and visitors |
80+Principal Investigators |
£150Mworth of active grants |
|---|
Our strategic priorities
The Department’s scientific strategic priorities are focused around the following 4 sections:
Cardiovascular and Respiratory Medicine (CaRM)
CaRM covers all research into cardiovascular disease, respiratory conditions and related topics in Cambridge. CaRM PIs and Associate PIs are mostly based in VPD-HLRI, working closely with clinicians at Addenbrooke’s and Papworth hospitals and extending into Cambridge Cardiovascular.
In 2024, CaRM successfully renewed its status as a British Heart Foundation Centre for Research Excellence. In collaboration with the Cardiorespiratory BRC theme, CaRM aims to improve cardiovascular and lung health across the world by working collaboratively across the university, NHS, commercial and charitable sectors.
Section lead: Professor Martin Bennett
Immunology and Infectious Disease (ImID)
IMID unites expert groups in immune-mediated and infectious diseases. Researchers at IMID are based at Addenbrooke’s Hospital, CITIID, and the MRC Laboratory of Molecular Biology. IMID also launched the Cambridge Immunology Network to connect and collaborate with immunology researchers across Cambridge and beyond.
A major focus of ImID is fostering global health research through partnerships with institutions such as National University of Singapore - Cambridge Immune Phenotyping Centre, the Hong Kong Jockey Club Global Health Institute.
Section lead: Professor Menna Clatworthy
Perioperative, Acute, Critical Care and Emergency Medicine (PACE)
PACE makes substantial clinical and research contributions, with our clinical academics holding leading roles in national policy and clinical trials. Notably, PACE leads on the Lancet Neurology Commission on Traumatic Brain Injury and coordinating the International Initiative on Traumatic Brain Injury Research.
PACE PIs and affiliated PIs are based at the Addenbrooke’s Hospital and VPD-HLRI.
Section lead: Professor Jon P Coles
Specialty Medicine, Research and Training (SMaRT)
SMaRT oversees training and research connections between the Department of Medicine and the NHS through formal links with NHS Clinical Service Leads in both the Cambridge University & Royal Papworth Hospitals. SMaRT also coordinates clinical medical student engagement and clinical academic trainings, advises affiliated postholders, and hosts the weekly medical Grand Round to promote closer ties between members of the Department and NHS colleagues.
All Department of Medicine PIs and Associate PIs are also Members of SMaRT.